Verifiable federated learning for medical AI — patient data never leaves the hospital, every computation is cryptographically proven on 0G Chain.

NeuroLedger solves the core problem in medical AI: hospitals cannot legally share patient data, so AI models train in isolation and stay mediocre. We built a federated learning network where hospitals collaborate on AI training without sharing any patient data.
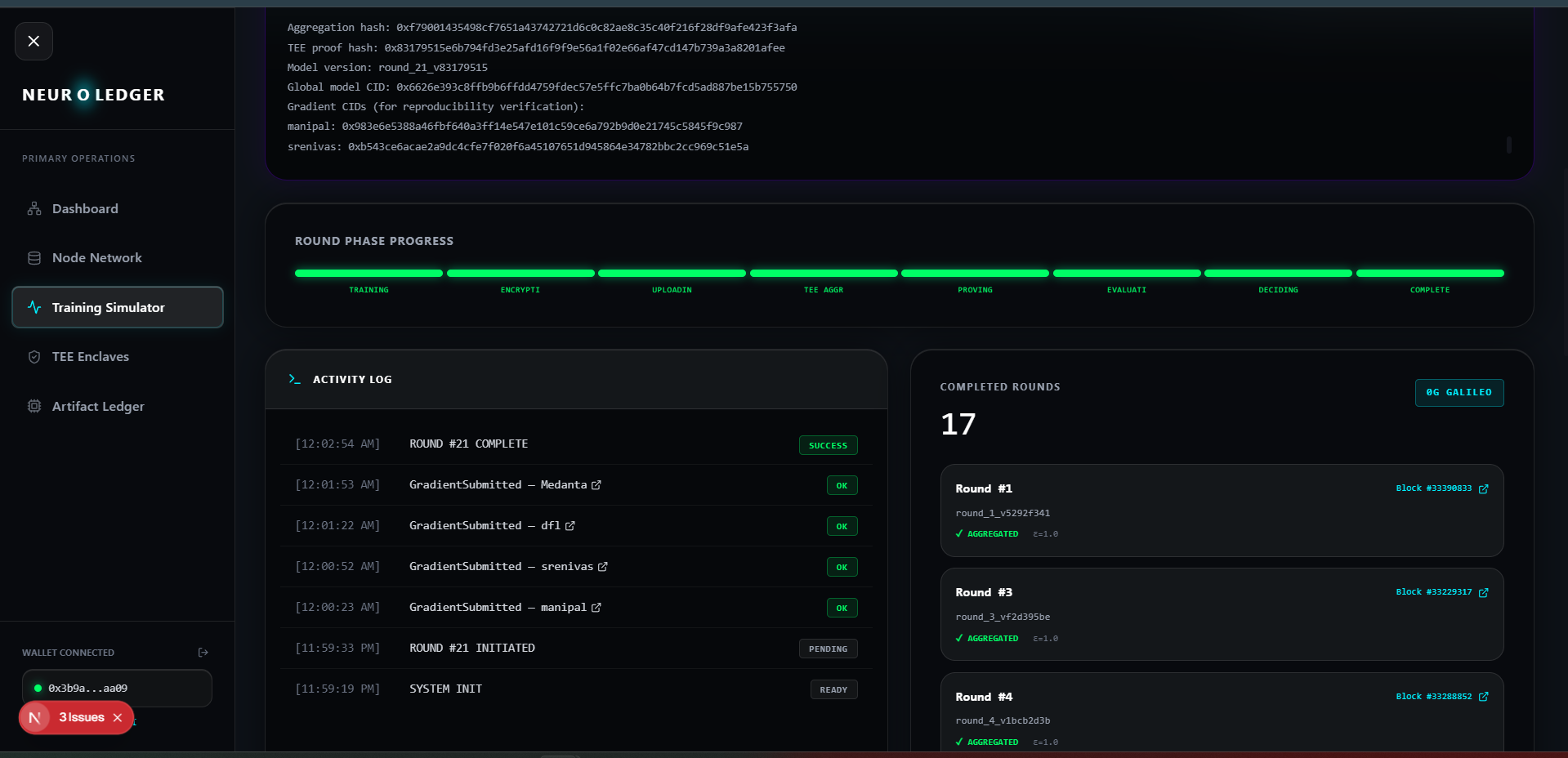
Each hospital agent uploads their patient CSV, trains a logistic regression model locally, and applies Rényi Differential Privacy noise (ε=1.0) to the gradient before it leaves the machine. The DP-noised gradient is uploaded to 0G Storage (turbo network) as a content-addressed file. Its Merkle root hash and CID are submitted to the NeuroLedger smart contract on 0G Chain as a GradientSubmitted event.
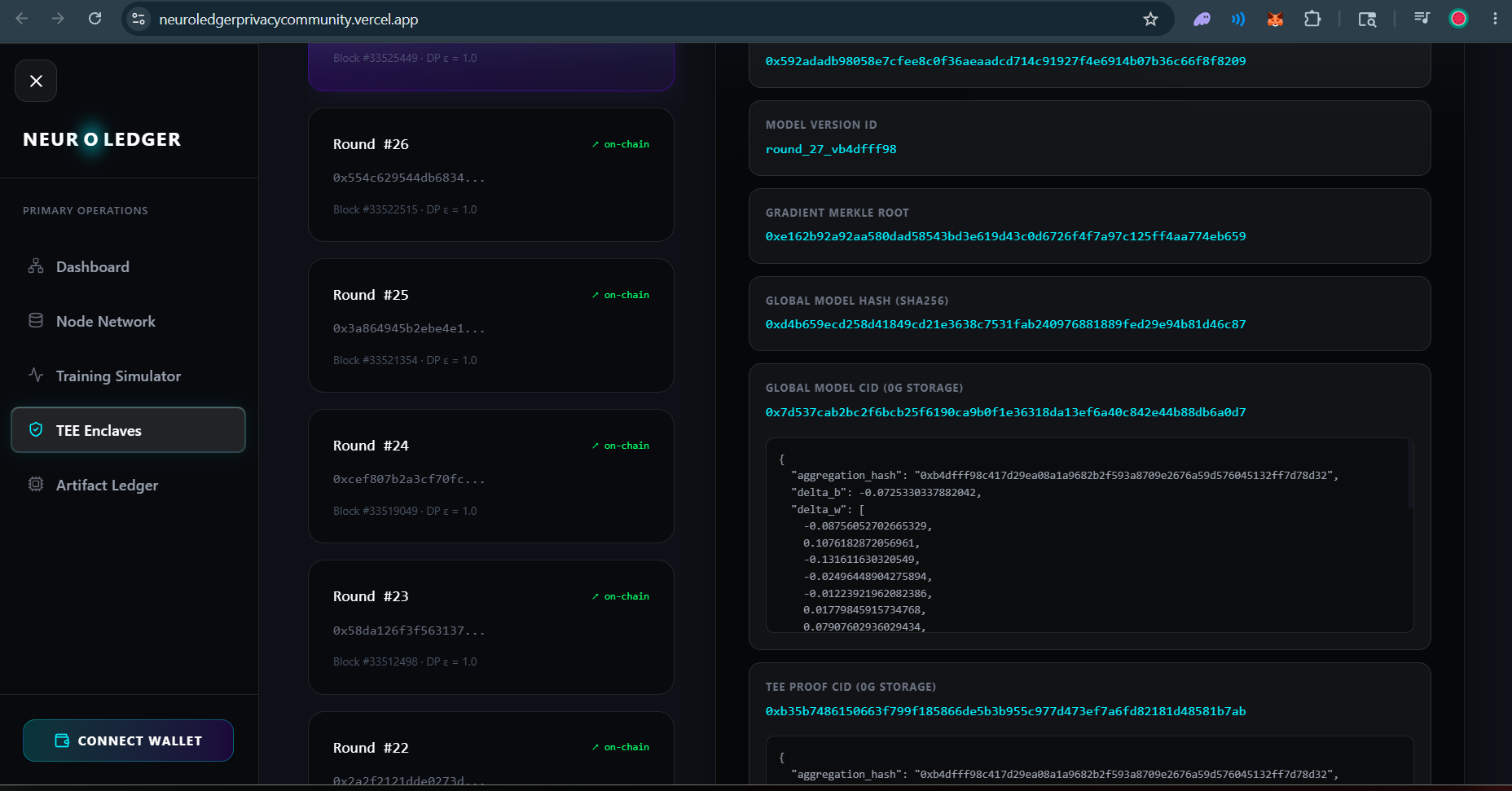
The aggregation runs inside an Intel TDX hardware enclave via 0G Compute's TeeML network. Multi-Krum and Trimmed Mean algorithms provide Byzantine robustness. The enclave produces a real hardware attestation (mr_enclave, mr_signer non-zero, inference_valid=True). This attestation JSON is stored on 0G Storage and its root hash is anchored on-chain in the AggregationComplete event.
The defining feature: anyone can download the gradient CIDs from 0G Storage and reproduce the exact aggregation hash on-chain using our verification script. The computation is deterministically verifiable by any third party — not just claimed.
Built and deployed during the hackathon:
- NeuroLedger.sol with 8-event lifecycle on 0G Galileo Testnet + 0G Aristotle Mainnet
- 21+ completed rounds with real UCI medical datasets
- 6 hospitals as on-chain agents with staked AOGI
- Real TeeML hardware attestation every round since round 18
- Hospital CSV upload with automatic data deletion after gradient computation
- Full Next.js frontend deployed on Vercel with live chain data
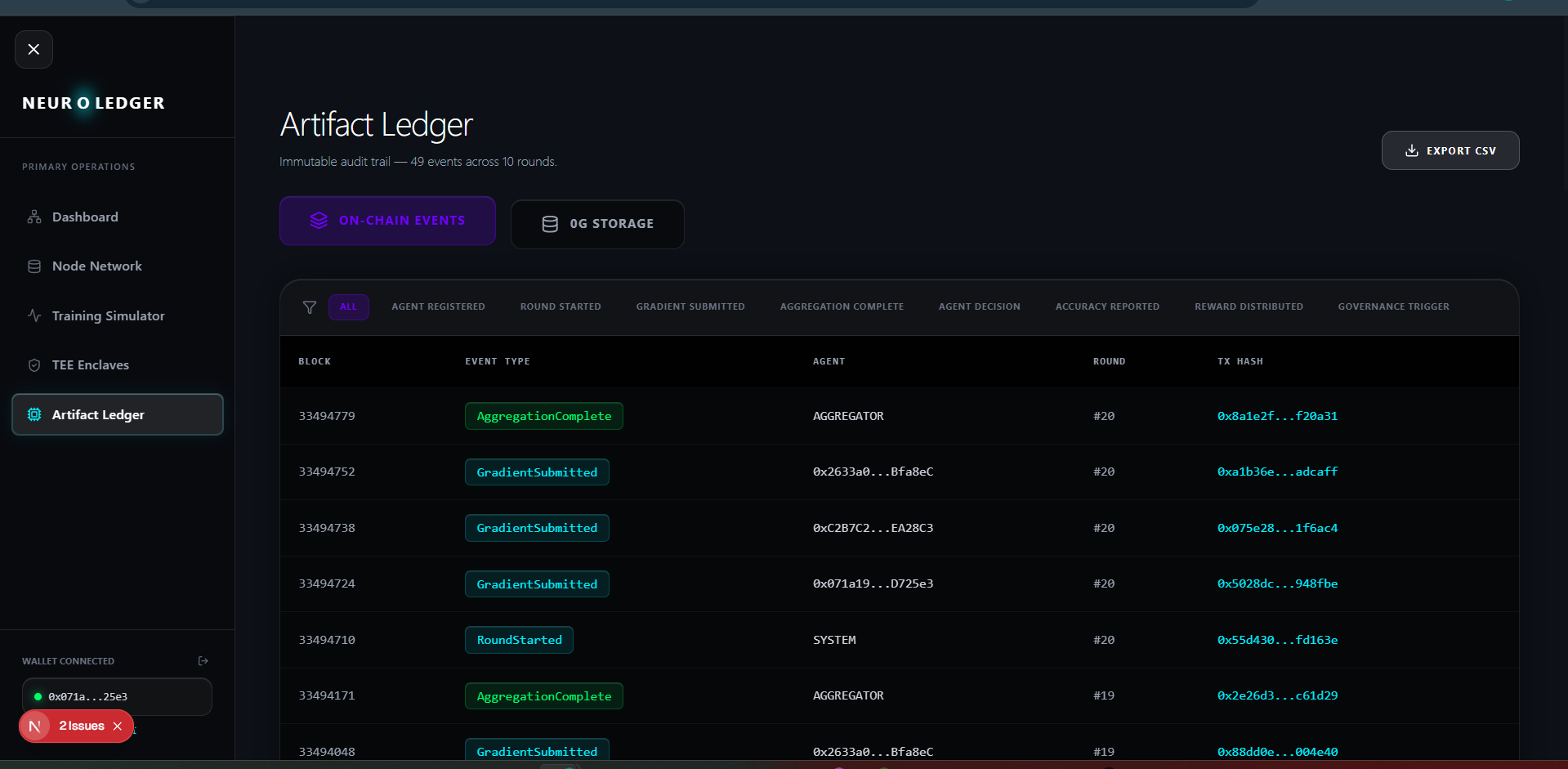
- 49+ on-chain events auditable on chainscan-galileo.0g.ai
0G components used: 0G Chain (smart contract), 0G Storage turbo (gradient and model files), 0G Compute TeeML (hardware-attested aggregation).
Week 1 — Core Infrastructure:
Designed architecture around 0G's full stack. Built NeuroLedger.sol with
8-event lifecycle. Deployed to 0G Galileo. Completed first end-to-end round
with 3 hospitals submitting real gradients on-chain. Fixed gas limit issues,
startRound event parsing, and aggregator role management.
Week 2 — Real Integrations:
Switched 0G Storage from standard (down) to turbo network using the official
TypeScript SDK — real Merkle root CIDs replacing SHA256 fallbacks. Integrated
0G Compute TeeML via compute starter kit. Achieved first round with real Intel
TDX hardware attestation (inference_valid=True). Built SSE streaming for live
runner output. Implemented hospital CSV upload with DP privacy deletion.
Week 3 — Scale + Polish:
Built full Next.js frontend: Training Simulator with phase progress, TEE
Enclaves panel, Artifact Ledger with 49+ events, dynamic hospital selector
reading from chain, dataset upload with class distribution stats. Registered
6 hospitals on-chain. Ran 21+ rounds. Deployed to Vercel. Deployed contract
to 0G Aristotle Mainnet. Built reproducibility verification script.
Final numbers: 21+ rounds, 6 hospitals, 49+ events, real TeeML every round.
Not currently fundraising.
NeuroLedger is a working prototype with real on-chain proof. The next step
is piloting with actual hospitals in APAC where health data regulations are
actively evolving (Singapore PDPA, India DPDP Act 2023). Open to conversations
with healthcare institutions, research hospitals, or infrastructure teams
interested in verifiable medical AI.